Table of Contents (click to expand)
Initiation, elongation and termination are three main steps in DNA replication. Let us now look into more detail of each of them:
Step 1: Initiation
The point at which the replication begins is known as the Origin of Replication (oriC). Helicase brings about the procedure of strand separation, which leads to the formation of the replication fork.
Step 2: Elongation
The enzyme DNA Polymerase III makes the new strand by reading the nucleotides on the template strand and specifically adding one nucleotide after the other. If it reads an Adenine (A) on the template, it will only add a Thymine (T).
Step 3: Termination
When Polymerase III is adding nucleotides to the lagging strand and creating Okazaki fragments, it at times leaves a gap or two between the fragments. These gaps are filled by ligase. It also closes nicks in double-stranded DNA.
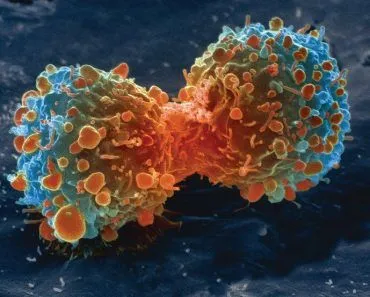
We all know that each human being begins their life as a single cell, which divides to form two cells, and these two go on to form four! This process helps us to form our tiny little body, which then grows into an adult! Now while all this is happening, our DNA is also being divided into these cells. But does the cell divide the existing DNA into two parts? Or does it make a second copy? If you think it is the latter, then you are correct! The cell does make a second copy, so when two daughter cells are formed; each one of them gets a complete set of DNA.
Recommended Video for you:
Structure Of DNA
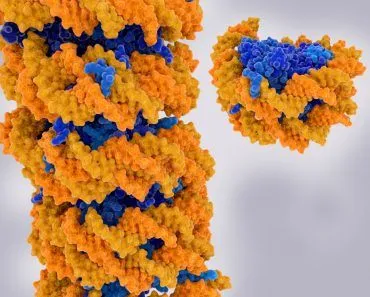
Before we jump into the process of replication, let us take a quick look at the structure of DNA.
As we all know, DNA is the genetic code that helps our cells to develop and reproduce in a planned way. Because of which it is called the ‘Blueprint of Life’.
DNA is the genetic material that defines cells in bodies. In order for a cell to duplicate and divide into its daughter cells (either through the process of meiosis or mitosis), organelles and biomolecules must be copied first and then distributed among all cells.
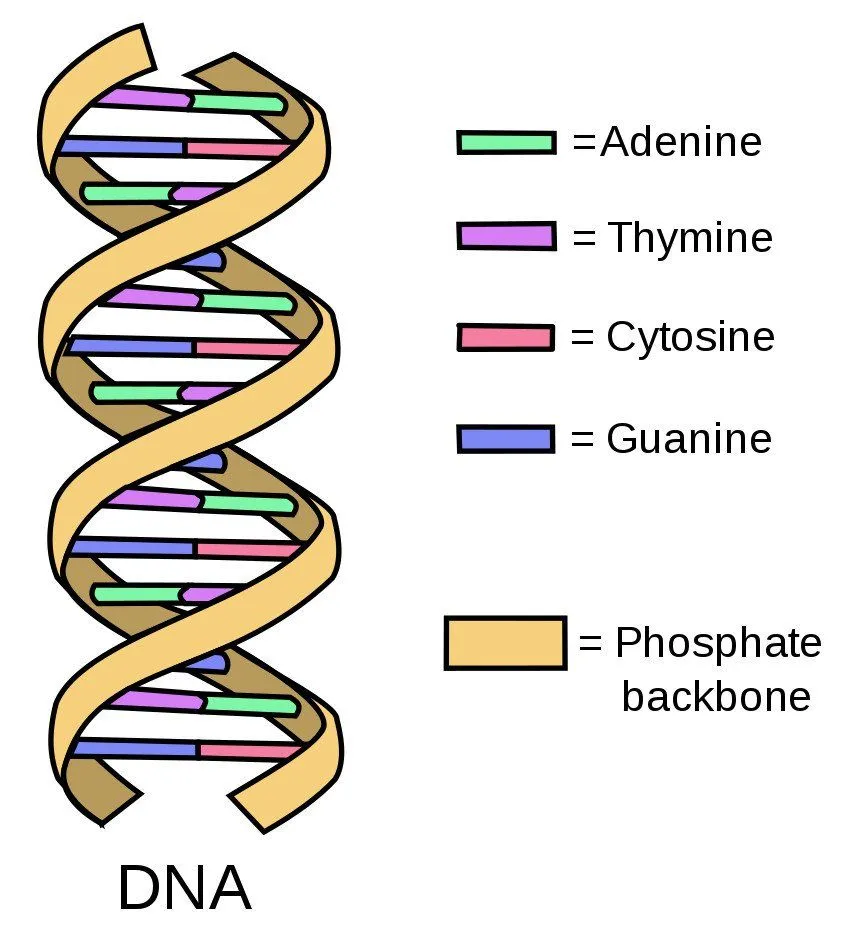
Getting back to its structure, DNA is made up of four nucleotides. Thinking what Nucleotides are? They are molecules, which are made of a phosphate group, a sugar ring, and a nitrogen base! These nucleotides are Adenine (A), Thymine (T), Guanine (G), and Cytosine (C). A and G are called Purines while T and C are called Pyrimidines. Those words can be a mouthful but you will be able to read them after a little bit of practice.
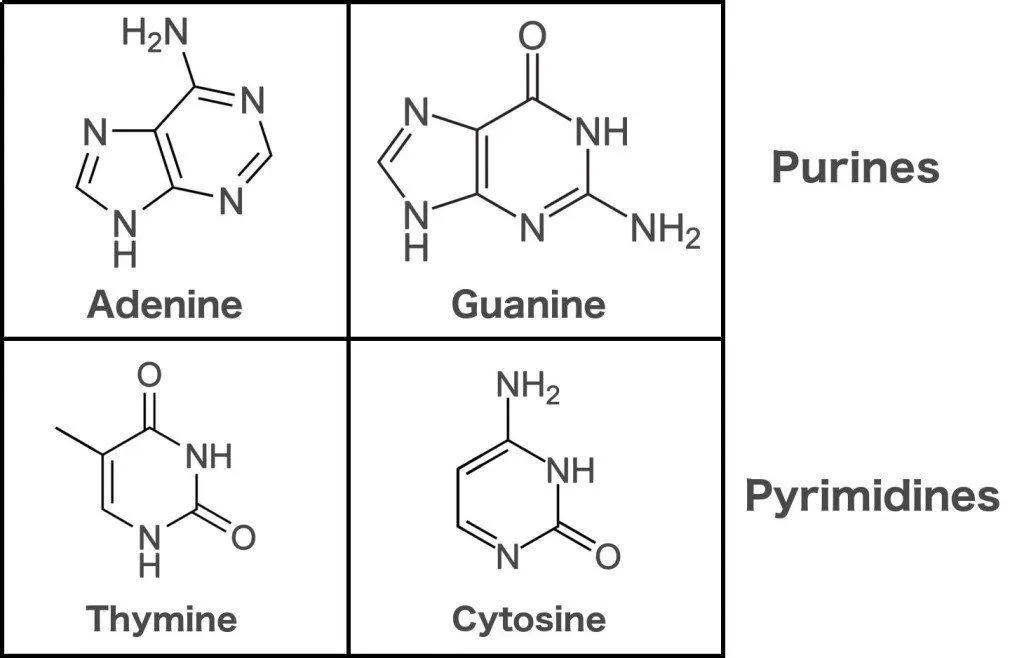
DNA is made of two strands. These strands have nucleotides lined up one after the other and those nucleotides are bound to the nucleotides on the other strand to create a ladder-like structure! Now the binding between nucleotides is very specific and the binding is via Hydrogen Bonds. A will bind to T and C will bind to G. These nucleotides bind to each other and are called as Base pairs. So there we have it. A seemingly never-ending ladder made of nucleotides pairing up with each other. But there is one more change, take that ladder and twist it! That’s it, our DNA looks like a simple double helix with specific nucleotide binding. Easy, right?
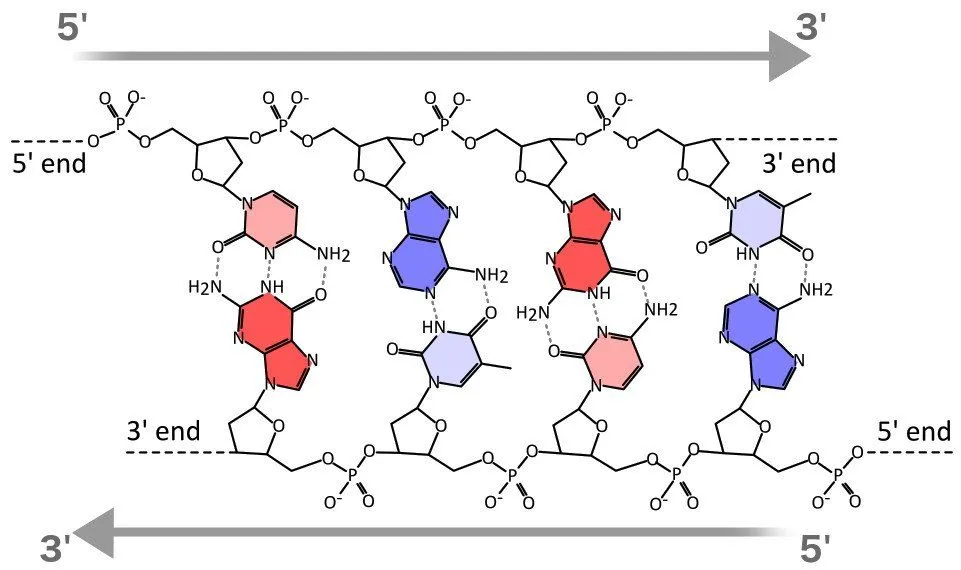
Directionality
These strands have two designated ends called 5’ and 3’ (you can read that as 5 prime end and 3 prime end). These numbers indicate end-to-end chemical orientation. The numbers 5 and 3 represent the fifth and third carbon atom of the sugar ring respectively. 5’ is the end, which joins a phosphate group that attaches to another nucleotide. 3’ end is important as during replication the new nucleotide is added to this end.
In terms of direction, if one strand is 5’ to 3’ while reading from left to right, the other strand will be 3’ to 5’. Simply put, the strands run in opposite directions. This orientation is kept for easy binding between nucleotides of the opposite strands.
Process Of Replication
Replicating the entire DNA is no easy job. The human genome (Genome means a complete set of genes present in the cell) has around 3 billion base pairs (Nucleotide pairing, remember?). So to make a copy of something that long would take a lot of time. But it doesn’t! Because our cells have a set of enzymes and proteins which makes this process quick!
Each enzyme and protein have their own specific function. Let us look at the process step by step.
Initiation
- Helicase – The point at which the replication begins is known as the Origin of Replication. Helicase brings about the procedure of strand separation, which leads to the formation of the replication fork. It breaks the hydrogen bond between the base pairs to separate the strand. It uses energy obtained from ATP Hydrolysis to perform the function.
- SSB Protein – Next step is for the Single-Stranded DNA Binding Protein to bind to the single-stranded DNA. Its job is to stop the strands from binding again.
- DNA Primase – Once the strands are separated and ready, replication can be initiated. For this, a primer is required to bind at the Origin. Primers are short sequences of RNA, around 10 nucleotides in length. Primase synthesizes the primers.
Elongation
- DNA Polymerase III – This enzyme makes the new strand by reading the nucleotides on the template strand and specifically adding one nucleotide after the other. If it reads an Adenine (A) on the template, it will only add a Thymine (T). It can only synthesize new strands in the direction of 5’ to 3’. It also helps in proofreading and repairing the new strand. Now you might think why does Polymerase keep working along the strand and not randomly float away? Its because a ring-shaped protein called as sliding clamp holds the polymerase into position.
Now when replication fork moves ahead and the Polymerase III starts to synthesize the new strand a small problem arises. If you remember, I mentioned that the two strands run in the opposite directions. This means that when both strands are being synthesized in 5’ to 3’ direction, one will be moving in the direction of the replication fork while the other will move in the opposite.
The strand, which is synthesized in the same direction as the replication fork, is known as the ‘leading’ strand. The template for this strand runs in the direction of 3’ to 5’. The Polymerase has to attach only once and it can continue its work as the replication fork moves forward. However, for the strand being synthesized in the other direction, which is known as the ‘lagging’ strand, the polymerase has to synthesize one fragment of DNA. Then as the replication fork moves ahead, it has to come and reattach to the new DNA available and then create the next fragment. These fragments are known as Okazaki fragments (named after the scientist Reiji Okazaki who discovered them).
Termination
- DNA Polymerase I – If you remember, we had added a RNA primer at the Origin to help Polymerase initiate the process. Now as the strand has been made, we need to remove the primer. This is when Polymerase I comes into the picture. It takes the help of RNase H to remove the primer and fill in the gaps.
- DNA ligase – When Polymerase III is adding nucleotides to the lagging strand and creating Okazaki fragments, it at times leaves a gap or two between the fragments. These gaps are filled by ligase. It also closes nicks in double-stranded DNA.
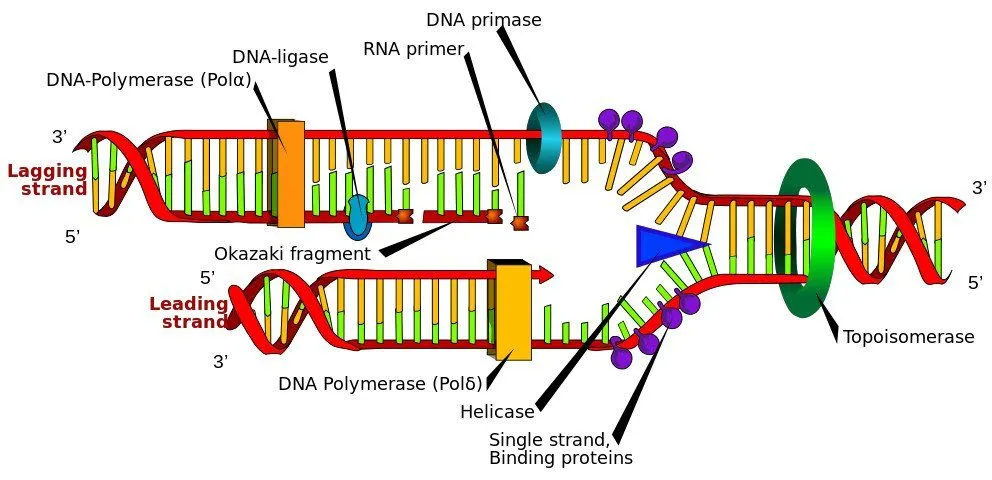
The Replication process is finally complete once all the primers are removed and Ligase has filled in all the remaining gaps. This process gives us two identical sets of genes, which will then be passed on to two daughter cells. Every cell completes the entire process in just one hour!
The reason for taking such short amount of time is multiple Origins. The cell initiates the process from a number of points and then the pieces are joined together to create the entire genome!
It is important for DNA present in the nucleus to be replicated so that every new cell receives the appropriate number of chromosomes. Overall, this process is crucial for cell repair and growth and reproduction in living organisms.